-Search query
-Search result
Showing 1 - 50 of 52 items for (author: cianfrocco & ma)

EMDB-41621: 
Full-length P-Rex1 in complex with inositol 1,3,4,5-tetrakisphosphate (IP4)
Method: single particle / : Cash JN, Tesmer JJG

EMDB-24672: 
The cryo-EM map of KIF18A bound to KIFBP
Method: single particle / : Tan Z, Solon AL, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

EMDB-24677: 
Cryo-EM structure of KIFBP core
Method: single particle / : Solon AL, Tan Z, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

EMDB-24744: 
Cryo-EM structure of KIFBP:KIF15
Method: single particle / : Solon AL, Tan Z, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

EMDB-24745: 
Cryo-EM map of KIFBP
Method: single particle / : Tan Z, Solon AL, Cianfrocco MA

PDB-7rsi: 
The cryo-EM map of KIF18A bound to KIFBP
Method: single particle / : Tan Z, Solon AL, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

PDB-7rsq: 
Cryo-EM structure of KIFBP core
Method: single particle / : Solon AL, Tan Z, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

PDB-7ryp: 
Cryo-EM structure of KIFBP:KIF15
Method: single particle / : Solon AL, Tan Z, Schutt KL, Jepsen L, Haynes SE, Nesvizhskii AI, Sept D, Stumpff J, Ohi R, Cianfrocco MA

PDB-7ryq: 
Cryo-EM map of KIFBP
Method: single particle / : Tan Z, Solon AL, Cianfrocco MA

EMDB-21692: 
PKA RIIbeta holoenzyme with DnaJB1-PKAc fusion in fibrolamellar hepatoceullar carcinoma
Method: single particle / : Lu TW, Aoto PC

EMDB-21693: 
PKA RIIbeta holoenzyme with DnaJB1-PKAc fusion in fibrolamellar hepatoceullar carcinoma
Method: single particle / : Lu TW, Aoto PC

EMDB-22754: 
Aldolase, rabbit muscle (no beam-tilt refinement)
Method: single particle / : Cianfrocco MA, Kearns SE

EMDB-22755: 
Aldolase, rabbit muscle (beam-tilt refinement x1)
Method: single particle / : Kearns SK, Cash JN

EMDB-22756: 
Aldolase, rabbit muscle (beam-tilt refinement x2)
Method: single particle / : Kearns SK, Cash JN

EMDB-22757: 
Aldolase, rabbit muscle (beam-tilt refinement x3)
Method: single particle / : Kearns SK, Cash JN

EMDB-22758: 
Aldolase, rabbit muscle (beam-tilt refinement x4)
Method: single particle / : Kearns SK, Cash JN

EMDB-21490: 
Influenza hemagglutinin trimer preprocessed with automatic workflow
Method: single particle / : Li Y, Cianfrocco MA

EMDB-21491: 
Thermoplasma acidophilum 20S proteasome cryo-EM structure using automatic preprocessing workflow
Method: single particle / : Cianfrocco MA, Li Y

EMDB-21492: 
Rabbit muscle aldolase cryo-EM structure using automatic preprocessing workflow
Method: single particle / : Cianfrocco MA, Li Y

EMDB-20512: 
Cryo-EM structure of MLL1 in complex with RbBP5 and WDR5 bound to the nucleosome
Method: single particle / : Park SH, Ayoub A

EMDB-20513: 
Cryo-EM structure of MLL1 in complex with RbBP5 and WDR5 bound to the nucleosome
Method: single particle / : Park SH, Ayoub A

EMDB-20514: 
Cryo-EM structure of RbBP5 bound to the nucleosome
Method: single particle / : Park SH, Ayoub A

EMDB-20308: 
Single Particle Reconstruction of Phosphatidylinositol (3,4,5) trisphosphate-dependent Rac exchanger 1 bound to G protein beta gamma subunits
Method: single particle / : Cash JN, Cianfrocco MA

EMDB-8908: 
Cryo-EM structure of beta-galactosidase using RELION on Amazon Web Services
Method: single particle / : Cianfrocco MA, Lahiri I, DiMaio F, Leschziner AE

PDB-6drv: 
Beta-galactosidase
Method: single particle / : Cianfrocco MA, Lahiri I, DiMaio F, Leschziner AE

EMDB-7038: 
RNA Pol II elongation complex bound to Rad26
Method: single particle / : Lahiri I, Leschziner AE

EMDB-8736: 
Structural Basis forEukaryotic Transcription-Coupled Repair Initiation
Method: single particle / : Lahiri I, Leschziner AE

EMDB-8885: 
RNA Pol II complex arrested by a CPD lesion
Method: single particle / : Lahiri I, Leschziner AE

EMDB-8735: 
Ternary complex of RNA Pol II, transcription scaffold and Rad26
Method: single particle / : Lahiri I, Leschziner AE

EMDB-8737: 
RNA pol II elongation complex
Method: single particle / : Lahiri I, Leschziner AE

PDB-5vvr: 
Ternary complex of RNA Pol II, transcription scaffold and Rad26
Method: single particle / : Lahiri I, Leschziner AE

PDB-5vvs: 
RNA pol II elongation complex
Method: single particle / : Lahiri I, Leschziner AE

EMDB-8673: 
Cryo-EM structure of yeast cytoplasmic dynein-1 with Lis1 and ATP
Method: single particle / : Cianfrocco MA, DeSantis ME, Htet ZM, Tran PT, Reck-Peterson SL, Leschziner AE

EMDB-8706: 
Cryo-EM structure of yeast cytoplasmic dynein with Walker B mutation at AAA3 in presence of ATP-VO4
Method: single particle / : Cianfrocco MA, DeSantis ME, Htet ZM, Tran PT, Reck-Peterson SL, Leschziner AE

PDB-5vh9: 
Cryo-EM structure of yeast cytoplasmic dynein-1 with Lis1 and ATP
Method: single particle / : Cianfrocco MA, DeSantis ME, Htet ZM, Tran PT, Reck-Peterson SL, Leschziner AE

PDB-5vlj: 
Cryo-EM structure of yeast cytoplasmic dynein with Walker B mutation at AAA3 in presence of ATP-VO4
Method: single particle / : Cianfrocco MA, DeSantis ME, Htet ZM, Tran PT, Reck-Peterson SL, Leschziner AE
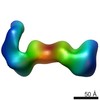
EMDB-6431: 
RCT reconstruction of CMV gHgLgO and pentameric complexes bound to highly neutralizing antibodies
Method: single particle / : Ciferri C, Cianfrocco MA, Chandramouli S, Carfi A
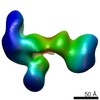
EMDB-6432: 
RCT reconstruction of glycoprotein gHgLgO in complex with neutralizing Fab fragments 3G16 and 13H11
Method: single particle / : Ciferri C, Cianfrocco MA, Chandramouli S, Carfi A
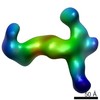
EMDB-6436: 
RCT reconstruction of CMV complex
Method: single particle / : Ciferri C, Cianfrocco MA, Chandramouli S, Carfi A
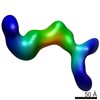
EMDB-6437: 
RCT reconstruction of CMV protein complex
Method: single particle / : Ciferri C, Cianfrocco MA, Chandramouli S, Carfi A

EMDB-6438: 
Structure of CMV protein complex
Method: single particle / : Ciferri C, Cianfrocco MA, Chandramouli S, Carfi A

EMDB-2858: 
Cryo-EM structure of yeast 80S ribosome calculated on Amazon's elastic cloud computing network
Method: single particle / : Cianfrocco MA, Leschziner AE

EMDB-2282: 
TFIIA-induced conformational changes of human TFIID suggest a modular architecture that is required for core promoter specific interactions
Method: single particle / : Cianfrocco MA, Grob P, Fang J, Kassavetis GA, Juven-Gershon T, Kadonaga JT, Nogales E
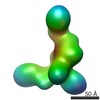
EMDB-5601: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW

EMDB-5602: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW

EMDB-5603: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW
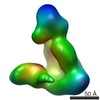
EMDB-5604: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW

EMDB-5605: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW
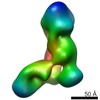
EMDB-5606: 
Substrate-specific structural rearrangements of human Dicer
Method: single particle / : Taylor DW, Ma E, Shigematsu H, Cianfrocco MA, Noland CL, Nagayama K, Nogales E, Doudna JA, Wang HW

EMDB-2287: 
Human TFIID binds to core promoter DNA in a reorganized structural state
Method: single particle / : Cianfrocco MA, Kassavetis GA, Grob P, Fang J, Juven-Gershon T, Kadonaga JT, Nogales E
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
